Biomarker2Validate: a pathway-driven customizable workflow for biomarker analysis
- Jun 25, 2023
- 4 min read
Updated: Jul 26, 2023
In recent years, scientists have been actively using -omics data for biomarker analysis by leveraging established and new technologies such as mass spectrometry, bulk RNA-seq, single-cell transcriptomics, CITE-seq, proteomics, epigenetics, and metabolomics. With this continuum of data, a plethora of bioinformatics methods have evolved over the past few years to help us make sense of the data, find important genes, proteins, and pathways - the insights from which can shed light on potential candidates for biomarker discovery.
So, what’s next?
Even when we have identified candidate biomarkers, validating them is a lengthy and arduous process. In addition to being sensitive and specific, an ideal biomarker should be detectable via accessible specimens (such as blood or urine). It needs to be either binary (i.e., present or absent) or quantifiable without subjective assessments. The result should be generated by an assay that can be applied in routine clinical practice and has a timely turnaround (i.e., in a matter of hours/days rather than weeks) (Ou et al., 2021). There are tools to support biomarker validation with such properties, but not for a comprehensive multi omics platform.
That’s why we developed Biomarker2Validate, a pathway-driven tunable and customizable workflow for biomarker analysis. The workflow enables identifying and assessing biomarkers across an established set of criteria. You can now try the Biomarker2Validate tool in the CDIAM Multi-Omics Studio. In this blog post, we will explain what Biomarker2Validate is and demonstrate the workflow with a breast cancer treatment response use case.

Biomarker2Validate methodology
Biomarker2Validate (B2V) is a workflow that predicts biomarkers via a novel biomarker weighting and prioritization algorithm, currently available in the CDIAM Multi Omics Studio. The workflow is adapted from the Pathway2Target workflow, developed by Pickett Lab at Brigham Young University, which is also incorporated in the CDIAM platform.
Biomarker2Validate takes a list of enriched pathways as inputs to identify potential biomarkers then applies a novel and customizable weighted scoring algorithm for biomarker prioritization. The weighted scores are computed using 5 key attributes listed below:
· Number of antibodies (default weight = 3)
· Whether the biomarker can be detected in blood, value is TRUE or FALSE (default weight = 3)
· Whether the biomarker can be detected in urine, value is TRUE or FALSE (default weight = 3)
· Number of significant pathways associated with the biomarker (default weight = 1)
· Number of associated diseases (default weight = 0.1)
Users can adjust the weights that these attributes should carry for prioritization. Biomarkers are eventually sorted in descending order by their weighted scores.
Biomarker2Validate has been incorporated into the CDIAM Multi Omics Studio, allowing users to input a gene expression matrix (from bulk RNAseq or single-cell RNAseq), a differentially expressed gene/protein list, or a gene/protein table and run biomarker analysis on such data.
Using Biomarker2Validate for breast cancer prognosis biomarker analysis
Data preparation
We retrieved bulk RNA-seq data from the study “Proline rich 11 (PRR11) overexpression amplifies PI3K signaling and promotes antiestrogen resistance in breast cancer” by Kyung-min Lee et al (2020). That data was collected from 36 responders and 22 non-responders to anti-estrogen treatment. These sequencing data were then mapped and quantified prior to using edgeR to identify differentially expressed (DE) genes. The DE results were loaded to CDIAM Multi Omics Studio to run enrichment. Finally, we performed Biomarker2Validate using the enrichment result to predict biomarkers.
Result
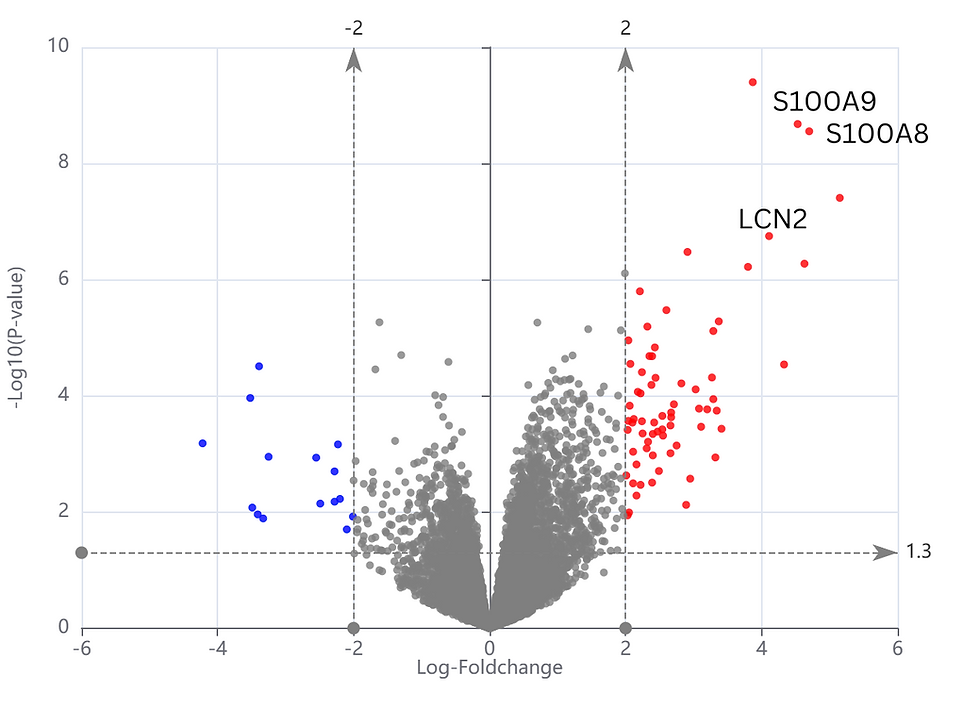
Biomarker2Validate results in CDIAM
From the Biomarker2Validate result, we identified LCN2, S100A8, S100A9 which satisfied the following criteria:
They exhibited high scores in the biomarker prioritization table (using Biomarker2Validate default set of weights);
They are detectable in blood and/or urine;
They were among the significant DE genes;
They were associated with at least one significant pathway in enrichment result;
There is at least one antibody for the biomarker
As suggested in the enrichment result in CDIAM, we also noticed that LCN2, S100A9, and S100A8 were involved in two significant pathways: neutrophil degranulation and metal sequestration by antimicrobial proteins. Aside from that, S100A9 and S100A8 were also associated with ER-Phagosome Pathway, MyD88 deficiency, IRAK4 deficiency and Regulation of TLR by endogenous ligand.

Interestingly, LCN2 has been known to facilitate tumorigenesis and promote the epithelial-to-mesenchymal transition (EMT). There was evidence of its association with aggressive types of breast cancers and poor prognosis (Leng et al., 2011, Xu et al, 2020), with the assumption that tumor cells exploit the beneficial innate immune function of LCN2 to support uncontrolled growth (Rodvold et al., 2012).
Meanwhile, S100A8 and S100A9 are members of S100 multigene subfamily of cytoplasmic EF-hand Ca+ binding proteins. They have also been suggested to be prognostic biomarkers for various cancer types such as Glioblastoma (Arora et al., 2019) and Non-small cell lung cancer (Kawai et al., 2011).
Through our Biomarker2Validate workflow, we identified LCN2, S100A9 and S100A8 to be potential breast cancer prognostic biomarkers for prioritization. Clearly, more investigation is needed and this brief analysis is for the purpose of demonstrating Biomarker2Validate capabilities to assist biomarker discovery and validation. If you are interested in having a deeper look into the result, contact us via support@pythiabio.com to get access to CDIAM and explore the Biomarker2Validate result through the link below:
>>> Access the Biomarker2Validate result by yourself:
References
Note: This article is for research use only.









Comments